Abstract
Introduction
Cordycepin is an antimicrobial substance that was first isolated from the entomopathogenic fungus Cordyceps militaris by Cunningham et al in 1950 (1). Since its discovery, a number of studies have focused on this substance (2,3). Cordycepin (Fig. 1A) with the molecular formula C10H13N5O3 has been demonstrated to have various bio-activities, including anticancer (4–7), antibacterial (8), antiviral (9,10) and immune regulation activities (11) and has became a topic of interest to various research groups.
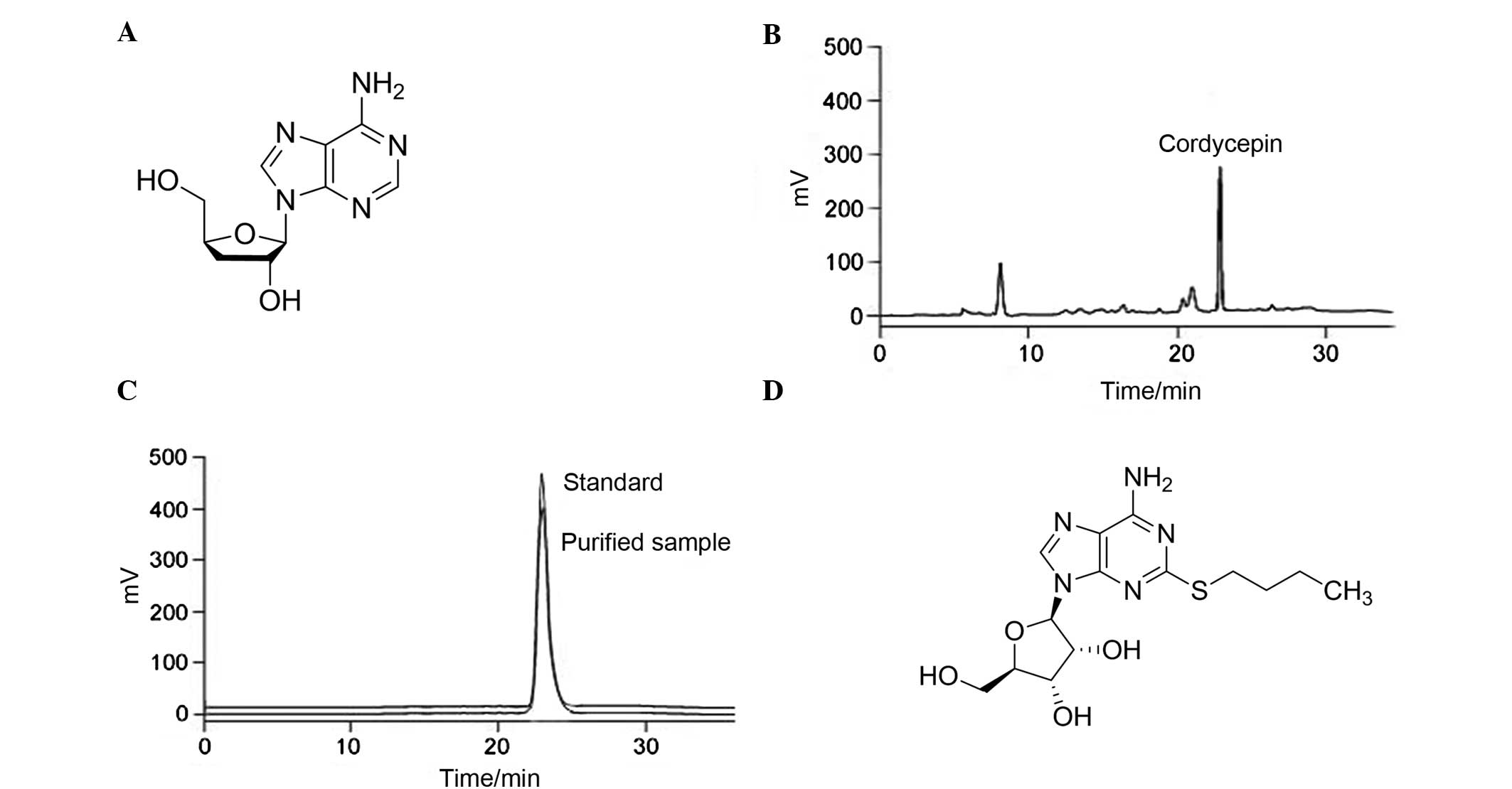 | Figure 1.Structure and HPLC chromatograms of cordycepin and the broad-spectrum adenosine-analog. (A) Structure of cordycepin. (B) HPLC chromatogram of crude cordycepin sample. (C) HPLC chromatogram of purified cordycepin sample compared with the standard. (D) Structure of the broad-spectrum adenosine-analog. HPLC, high-performance liquid chromatography. |
However, highly pure cordycepin is challenging to obtain; various drawbacks have been identified in the purification techniques of ion resin adsorption (12,13), silica gel column chromatography (14) and supercritical extraction (15). The high price of cordycepin acts as a significant barrier to the study of its bio-activity. Preparative high performance liquid chromatography (prep-HPLC) can shorten the production cycle, has relatively low operating costs, and is simple to carry out. Therefore, it is commonly used for the purification of chemical drugs and biological drugs by pharmaceutical companies and particularly in industrial purification (16). Prep-HPLC can be used to provide purified samples on a kilogram, or even ton, scale. Also, prep-HPLC is an effective technology currently used for natural product isolation in industry (17). The present study describes a new method of obtaining high-purity cordycepin by prep-HPLC, which may be useful for the large-scale isolation of cordycepin in the future.
Although studies have shown that cordycepin is a broad-spectrum antibiotic (18), studies focusing on the mechanism of the antibacterial activity and the possible targets are lacking. Molecular modeling and docking technology, which is efficient, accurate and simple to use, can reproduce the true interaction process between drug and targets that some of the current laboratory techniques are unable to detect, and is a method for predicting the mechanism of action of a drug (19). Therefore, molecular docking was also used in the present study to simulate the binding of cordycepin with different antibacterial targets, such as β-ketoacyl-(acyl-carrier-protein) synthase III (FabH), DNA gyrase B and thymidylate kinase, in order to identify the most likely antimicrobial mechanism of cordycepin. This should be of great valuable for future clinical and drug-resistance research.
Materials and methods
Materials
All chemicals and solvents used in this study were purchased from Sigma-Aldrich (St. Louis, MO, USA). The centrifuges (room-temperature, Allegra X-30; low temperature, GS-15R) were acquired from Beckman Coulter, Inc. (Brea, CA, USA). The biological purification columns were purchased from GE Healthcare Life Sciences (Chalfont, UK). The fermentation broths of Cordyceps militaris, Haemophilus influenzae (H. influenzae) KW20, Streptococcus pneumoniae (S. pneumoniae) R6, Escherichia coli (E. coli) MG1655 and Staphylococcus aureus (S. aureus) RN4200 were provided by Hangzhou Vocational and Technical College (Hangzhou, China). An analytical HPLC system (Waters 600) with a 2487 dual wavelength UV detector and analytical software (Waters Corporation, Milford, MA, USA) was used for the detection and analysis of cordycepin. A prep-HPLC system (Dalian Elite Analytical Instruments Co., Ltd., Dalian, China) with dual P280 constant pressure pump, UV200 II variable wavelength UV detector and analytical software was used for the purification of cordycepin. The 1H and 13C NMR spectra were recorded on a Bruker DPX 500 spectrometer (Bruker Corporation-Billerica, MA, USA). The interaction between cordycepin and different antibacterial targets was investigated using the CDOCKER protocol of Discovery Studio (version 3.1; Accelrys, Ltd., Cambridge, UK).
Purification of cordycepin by prep-HPLC
The fermentation broth of Cordyceps militaris with a cordycepin concentration of ~0.2 mg/ml was centrifuged at high speed (2,000 × g) for 15 min. The supernatant was concentrated under vacuum to a concentration of 0.6 mg/ml. Then, 1,800 ml of this sample solution was passed through 300 ml macroporous resin (HPD-100) for the preliminary removal of impurities. The volume flow rate of the loading sample was 2 BV/h, and the sample was eluted with ethanol at volume fraction of 25% at a volume flow of 3 bed volumes/h. The eluted sample was concentrated and freeze-dried to obtain the crude extract samples of cordycepin. The cordycepin contents of the crude samples were determined by HPLC. A solution was prepared by dissolving 400 mg crude sample in 100 ml mobile phase, and ultrasonic vibration was used to ensure that the sample was completely dissolved. This solution was subjected to further purification.
HPLC conditions were as follows: Column, symmetry shield C18 (250×4.6 mm; 5 µm); mobile phase, methanol:water (15:85, v/v); flow rate, 0.8 ml/min; UV detection, 260 nm; and injection quantity, 10 µl. The sample was filtered through a 0.45- µm membrane filter prior to injection. Quantitative analysis of cordycepin was conducted by evaluating the peak area on the basis of a standard curve. Peaks of cordycepin and other compounds in the sample were identified by their retention times and co-injection tests with corresponding standard compounds.
Prep-HPLC conditions were as follows: Column, Sinochrom ODS-BP (250×10 mm); mobile phase, methanol:water (15:85, v/v); flow rate, 1.0 ml/min; UV detection, 260 nm. A standard solution of cordycepin (1 ml) was injected into the prep-HPLC apparatus. The flow rate was 10 ml/min for observing the peak shape and retention time initially. Then, crude cordycepin solution (2 ml) was injected, with adjustment of the flow rate for complete separation and shortening of the sample peak time after observing the spectrum for crude cordycepin solution. The optimal flow rate was then determined and chromatographic fractions containing cordycepin were collected. The solvent was removed by freeze-drying. High purity cordycepin was obtained in crystalline form. In addition, the purified cordycepin was examined using NMR for comparison with the standard. These crystals were used for evaluating the inhibitory activity of cordycepin against various targets in the following experiments.
Experimental protocol of the docking study
In the present study, the binding mode of the original ligand (4-(2-amino-1, 3-thiazol-4-yl)pyrimidin-2-amine) and cordycepin were studied using molecular docking. Molecular docking of cordycepin into the three dimensional X-ray structure of NAD+-dependent DNA ligase (LigA) and other potential antibacterial targets was carried out using Discovery Studio, using the DS-CDOCKER protocol. The crystal structures, including that of LigA [Protein Data Bank (PDB) code: 3PN1] complex, were retrieved from the Research Collaboratory for Structural Bioinformatics PDB (http://www.rcsb.org/pdb/home/home.do). All bound water molecules and ligands were eliminated from the protein and polar hydrogens were added to the protein. Images showing the binding were prepared using Pymol software version 8.2.0.tar.bz2 (Slashdot Media Inc., San Diego, CA, USA).
In vitro experiments for the determination of IC50 values against LigA
Purification of DNA ligases from H. influenzae, S. pneumoniae, E. coli and S. aureus was conducted by the same procedure. In brief, 100 ml lysis buffer (Tiangen Biotech (Beijing) Co., Ltd., Beijing, China) was used to suspend the frozen cell paste (~1×106 cells). The cells were disrupted using a French press (model 1548-294US; Bodum USA Inc., New York, NY, USA) at 18,000 psi at 4°C, and the crude extract was centrifuged at 12,000 × g for 30 min at 4°C. A 20-ml Q-Sepharose HP (HR 16/10) column was pre-equilibrated with Buffer A [50 mM Tris-HCl, pH 7.5, 5 mM dithiothreitol (DTT), 2 mM ethylenediamine tetra-acetic acid (EDTA), 10% glycerol], and then the supernatant was added at a rate of 1.5 ml/min. After washing the column with Buffer A, the protein was eluted with a linear gradient (0 to 1 M NaCl in Buffer A). The ligase-containing eluents were combined, and dialyzed against 2 l Buffer A at 4°C overnight. The dialyzed sample was purified using a 20-ml Heparin Sepharose CL-6B (HR 16/10) column as described above. The fractions were pooled, and proteins were purified by the addition of 0.4 g/ml (NH4)2SO4. The mixture was kept on ice for 1 h. The sample was centrifuged at 5,000 × g for 30 min at 4°C. The sample was dispersed in 8 ml Buffer A, and loaded onto 320-ml Sephacryl S200 (HR 26/60) at a rate of 1 ml/min, which was pre-equilibrated with Buffer B (50 mM Tris/HCl, pH 7.5, 1 mM EDTA, 5 mM DTT, 10% glycerol, 150 mM NaCl). The ligase-containing eluent was pooled and dialyzed against 1 l Storage Buffer (50 mM Tris/HCl, pH 7.5, 1 mM EDTA, 100 mM KCl, 2 mM DTT, 20% glycerol) overnight at 4°C. The protein was characterized by sodium dodecyl sulfate-polyacrylamide gel electrophoresis and analytical liquid chromatography-mass spectrometry. The protein was stored at −80°C.
A fluorescence resonance energy transfer (FRET) assay was used for the analysis of LigA. The method was similar to that previously described (20,21). For the assays for H. influenzae LigA, 96-well black polystyrene flat-bottomed plates were used, and 100 µl reaction mixture (20% glycerol, 30 mM KCl, 30 mM (NH4)2SO4, 10 mM DTT, 1 mM EDTA, 0.002% Brij 35, 50 mM morpholinepropanesulfonic acid, 100 nM bovine serum albumin, 1 µM NAD+, 40 nM DNA substrate, 16 mM MgCl2 and 0.15 nM H. influenzae LigA) was added to the plates. The mixture was reacted at room temperature for 20 min, then 30 µl Quench reagent (8 M urea, 1 M Tris and 20 mM EDTA in water) was added as the reaction terminating agent. Plates were read using a Tecan Ultra plate reader (Tecan Group Ltd., Männedorf, Switzerland), with monitoring of the ratio of fluorescence intensities at two emission wavelengths (the TAMRA acceptor at 595 nm and the FAM donor at 535 nm) upon excitation of the FAM donor at 485 nm. The larger the ratio of 595 to 535 nm emission values, the larger the fraction of ligated DNA in the reaction. The ratio of the emission values at 595 and 535 nm was recorded. Control samples were also tested, which were 0.2% dimethyl sulfoxide (no compound) serving as the 0% inhibition control, and a mixture containing 50 mM EDTA as the 100% inhibition control. The inhibitory activity of the compound [cordycepin or a broad-spectrum adenosine analog (BCX4430; cat. no. HY-18649A; Medchemexpress LLC, Princeton, NJ, USA), used as a control] was measured at 10 different compound concentrations in the reaction system in order to determine the 50% inhibitory concentration (IC50). Assays for the other bacterial LigA isozymes were performed as described above for H. influenzae LigA, with the following changes in order to optimize enzyme performance, as previously described (22,23). S. aureus LigA reaction mixtures contained 0.15 nM enzyme, 72 nM DNA and 94 µM NAD+; S. pneumoniae LigA reaction mixtures contained 0.1 nM enzyme, 24 nM DNA and 6.8 µM NAD+; E. coli LigA reaction mixtures contained 0.1 nM enzyme, 22 nM DNA and 5.3 µM NAD+, with KCl and (NH4)2SO4 concentrations adjusted to 40 mM and 5 mM, respectively.
Results
HPLC analysis of cordycepin
Establishing the function between the mean peak area (Y) and the sample concentration (X) by HPLC, the linear regression equation was determined to be as follows: Y = 2.23 × 104X + 5,643 (R2=0.9998). Peak area and mass concentration exhibited a good linear relationship in the range of 0.112–56 µg/ml. The solution of crude cordycepin (0.1 ml) used for separation and preparation was diluted 100-fold for HPLC injection. The purity of the crude cordycepin sample was 12.1%, and the concentration of cordycepin following purification by prep-HPLC was 99.6%. The HPLC chromatograms of the crude sample and purified cordycepin are shown in Fig. 1B and C, respectively.
Components of cordycepin
The purified cordycepin was analyzed by NMR. The NMR results indicated that the purified cordycepin was consistent with the standard: 1H NMR (500 MHz, d6-DMSO): δ 8.34 [singlet (s), 1H], 8.13 (s, 1H), 7.26 (s, 2H), 5.86 [doublet (d), J=2.2 Hz, 1H], 5.64 (d, J=4.0 Hz, 1H), 5.14 [triplet (t), J=5.5 Hz, 1H], 4.57 (s, 1H), 4.34 [double doublet (dd), J=8.4, 6.1 Hz, 1H], 3.74-3.62 [multiplet (m), 1H], 3.56-3.45 (m, 1H), 2.32-2.18 (m, 1H), 1.91 (doublet of doubled of doublets, J=12.9, 6.2, 3.2 Hz, 1H); 13C NMR (125 MHz, d6-DMSO) δ 157.93, 154.31, 150.74, 140.97, 120.98, 92.69, 82.56, 76.46, 64.52, 35.96.
Docking analysis using the CDOCKER protocol
The CDOCKER protocol, which is used to precisely model molecular docking, was used to analyze the interaction between cordycepin and some common antibacterial targets. The results are shown in Table I.
Table I.Interaction between cordycepin and different proteins or kinases as determined by CDOCKER protocol. |
The interaction energy between cordycepin and LigA was the lowest, at −35.6055 kcal/mol, and thus it was hypothesized that cordycepin may achieve an antibacterial effect by binding to bacterial LigA. Therefore, the activity of cordycepin against different kinds of LigA was further tested and its activity was compared with that of a broad-spectrum adenosine analog (Fig. 1D) (20).
Biochemical potency of cordycepin
The activity results against LigA enzyme from S. pneumoniae, H. influenzae, E. coli and S. aureus are shown in Table II. The results show that the inhibitory activity of cordycepin against LigA is similar to that of the broad-spectrum adenosine analog. This may be because they have similar adenosine-based chemical structures.
Table II.Comparison of the biochemical potencies of cordycepin and broad-spectrum adenosine analog. |
Binding of compounds with LigA protein crystals
The binding modes of the ligand (4-(2-amino-1,3-thiazol-4-yl)pyrimidin-2-amine) and cordycepin were investigated using molecular docking. A comparison of the bonding of the two compounds with the LigA protein crystal is shown in Fig. 2. The results indicated that both compounds possess the same binding mode with LigA (PDB Code: 3PN1). Clearly, the two residues LEU-80 and SER-81 play a key role in the affinity of the small molecules with the target protein. The surface created by the residues within 4 Å close to the original ligand provide a deep binding pocket, and each of the two molecules can inserted into this cavity. These data indicate that cordycepin acts as a broad-spectrum antimicrobial agent by exerting an inhibitory effect against the LigA enzyme in different bacteria.
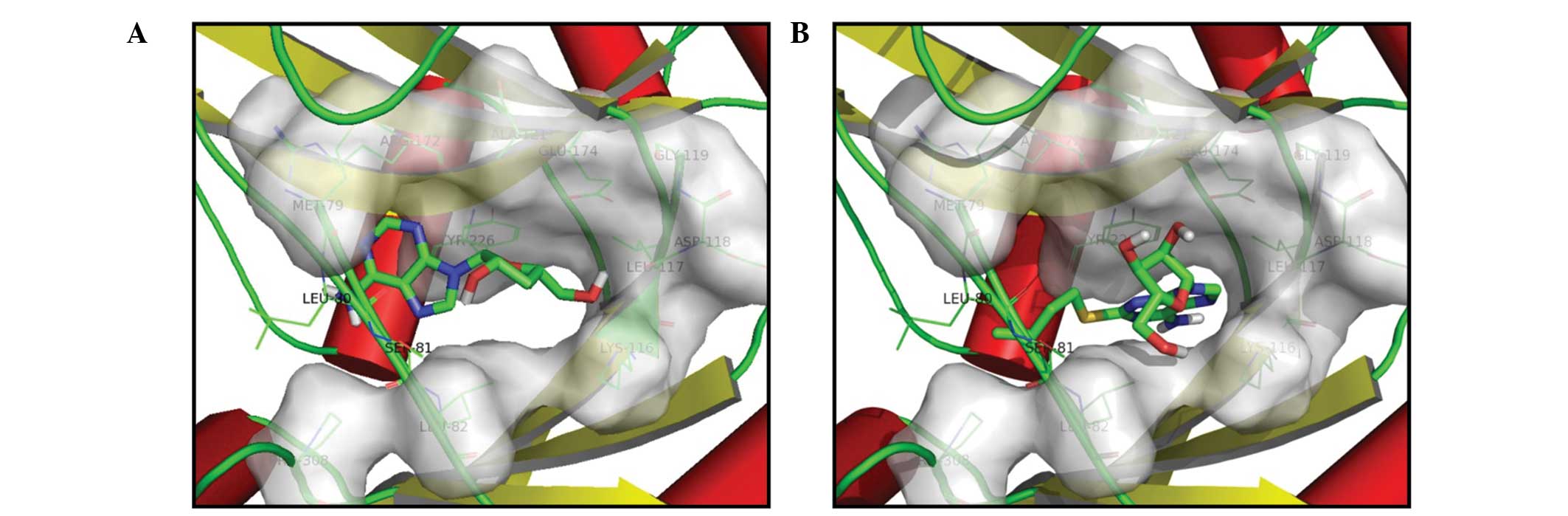 | Figure 2.Molecular docking of compounds into the three dimensional X-ray structure of Haemophilus influenzae NAD+-dependent DNA ligase. (A) Cordycepin; and (B) original ligand (4-(2-amino-1,3-thiazol-4-yl)pyrimidin-2-amine). |
Discussion
Cordycepin is a derivative of the nucleoside adenosine, which is a natural product originally extracted from fungi of the genus Cordyceps, but is now produced synthetically. Some enzymes are unable to discriminate between cordycepin and adenosine because of their similar structures. Therefore, cordycepin can participate in certain biochemical reactions; for example, it can be incorporated into RNA, thus causing its synthesis to be prematurely terminated (24–26).
In the present study, the antimicrobial activity and mechanism of action of cordycepin were explored. The mechanism of the antimicrobial activity of cordycepin has not been fully investigated previously and, to the best of our knowledge, a higher purity of cordycepin has not been obtained. A previous study indicated that prep-HPLC can shorten production cycles (16). In the present study, prep-HPLC was used to purify cordycepin, and the purity was up to 99.6%. Therefore, the results indicate that prep-HPLC is an important technique that may be useful for the large-scale isolation of cordycepin in the future.
Using a molecular docking method, the interaction energy between cordycepin and LigA was the lowest among all the potential targets that were analyzed. Therefore, it was speculated that cordycepin may achieve antibacterial effects by binding to bacterial LigA. To the best of our knowledge, this result has not been reported previously. The in vitro antibacterial assay for cordycepin indicated that cordycepin has similar inhibitory LigA activity to a broad-spectrum adenosine analog.
In summary, prep-HPLC was used to extract and purify cordycepin from the fermentation supernatant of Cordyceps militaris and the purity was up to 99.6%, which indicates that is may be useful for the large-scale isolation of cordycepin in the future. Also, molecular docking analysis was used to calculate the interaction energy of cordycepin with different antibacterial targets, and it was found that the interaction energy between cordycepin and LigA of H. influenzae was particularly low. To evaluate cordycepin as a LigA inhibitor, the inhibitory activity of cordycepin against LigA from H. influenzae, E. coli, S. pneumoniae and S. aureus was tested. Cordycepin showed inhibitory activity against LigA similar to that of a broad-spectrum adenosine analog. Therefore, cordycepin has the potential to be used as a broad-spectrum antibiotic with strong inhibitory activity against LigA.